Chemists have made a groundbreaking discovery in the field of enzyme research, as they have successfully determined the crystal structure of a crucial component of the monensin enzyme. This significant breakthrough has provided new insights into the mechanism of the enzyme’s reaction activity, offering potential opportunities for the development of safer and more effective antibiotics. Led by Professor Chu-Young Kim from the University of Illinois Urbana-Champaign (UIUC), the experimental team worked in collaboration with computational scientist Lela Vukovic from The University of Texas at El Paso (UTEP).
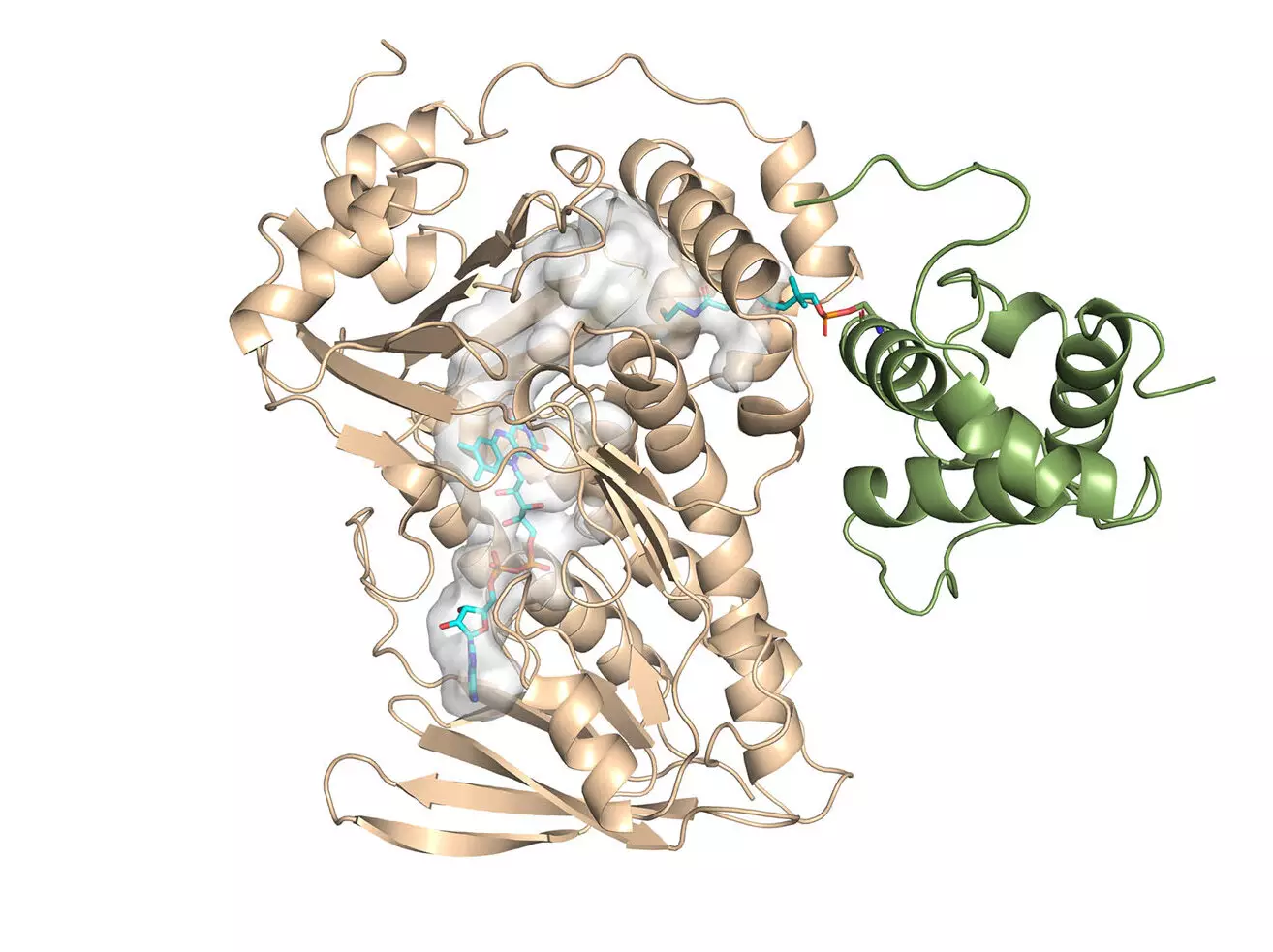
The main achievement of this study is the determination of the first crystal structure for a family of enzymes associated with monensin synthesis. MonCI, a key enzyme found in soil bacteria responsible for the natural production of monensin, was the focus of the research. Professor Kim and his colleagues successfully solved the crystal structure of MonCI, providing valuable insights into its composition and function. This groundbreaking discovery opens up new possibilities for the design and development of novel antibiotics.
One of the key findings of the study was the identification of the reaction sequence that produces monensin. The team discovered that MonCI carries out three crucial epoxidation reactions, a highly unusual process that has significant implications for engineering the bacterium to produce new antibiotics. Although the experimental approach was unable to obtain the crystal structure of the enzyme with the substrate in its active mode, computational modeling was used to simulate the stable structure and position of the substrate.
Lela Vukovic and her team from UTEP played a crucial role in the research by conducting extensive computational studies to analyze the stability and likely reactions of the substrate. Using the crystal structures determined by the experimentalists, Vukovic’s team developed comprehensive models of the enzyme-substrate system. This involved examining multiple systems and determining the most stable positions for the premonensin A and its epoxidated versions, which subsequently underwent the first, second, and third epoxidation reactions. The computational studies were made possible by utilizing supercomputers, such as TACC’s Lonestar6, which allowed for the characterization and understanding of these complex sequential reactions.
The research also paid tribute to the late Klaus Schulten, whose group developed the NAMD software that played a pivotal role in this study. The NAMD software provided the researchers with the ability to zoom into the enzyme and identify the order of reactions responsible for monensin production. The University of Illinois Urbana-Champaign continues to optimize the NAMD software for high-performance computing, ensuring its compatibility with the nation’s supercomputers. This ongoing development will enable future studies to explore and uncover the mechanisms of various enzymes.
The monensin biosynthesis pathway involves the coordinated action of at least 14 different enzymes, with MonCI being just one of them. While this study has shed light on the crystal structure and reaction mechanism of MonCI, there is still much to explore with the other enzymes involved in the process. As the researchers continue their investigation, they anticipate the development of improved versions of monensin that can better serve the healthcare needs of animals, particularly cattle and poultry. Furthermore, the toxic nature of monensin towards horses and dogs highlights the urgent need for a nontoxic alternative.
The determination of the crystal structure for a key component of the monensin enzyme has revolutionized our understanding of its reaction activity. This breakthrough not only provides valuable insights into the complex processes at play but also paves the way for the design and development of safer and more effective antibiotics. The collaboration between experimental and computational scientists has demonstrated the power of combining experimental data with computational modeling, unlocking new possibilities in enzyme research. As the journey continues, further investigations into the monensin biosynthesis pathway hold tremendous potential for improving animal healthcare and developing alternative treatments.


Leave a Reply