Researchers worldwide are working diligently to discover ways to combat bacteria that can resist current antibiotics to prevent a global health crisis. One potential target for the development of improved antibiotics is riboswitches, which are small RNA segments that regulate the production of proteins by bacterial cells. Riboswitches are almost exclusively found in bacteria, making them an ideal target for antibiotics that do not affect animals or humans. By gaining a complete understanding of how riboswitches function, scientists may be able to create drugs that disrupt the cellular machinery that creates necessary proteins.
New Findings Related to Riboswitches
Researchers from the University of Michigan’s Department of Chemistry and the Life Sciences Institute have discovered how a specific riboswitch regulates its own synthesis. The first step in creating a protein from the genetic code is called transcription, during which the RNA polymerase enzyme copies genetic information found in DNA into an RNA strand. RNAP undergoes several pauses during the process as it waits for further instructions from the cell. The mechanisms behind these pauses and restarts have long remained a mystery to scientists, but they provide a perfect target for antibiotics.
The team, led by Professor Nils Walter of the chemistry department, used biochemistry, structural biology, and computational modeling to determine how transcriptional regulation occurs. They looked at a particular riboswitch that binds a molecule made by the cell called preQ1. When the preQ1 molecule binds to the riboswitch, it alters the RNA’s shape, allowing RNAP to continue along the DNA so that transcription can continue.
Visualizing Riboswitches
Riboswitches were first discovered in 2002, but their specific roles in relation to transcription machinery have not been well understood. During their research, the Walter lab discovered that transcriptional pausing switches on and off as a result of the preQ1 molecule binding to the riboswitch. They teamed up with cryo-EM expert Professor Melanie Ohi to visualize the process.
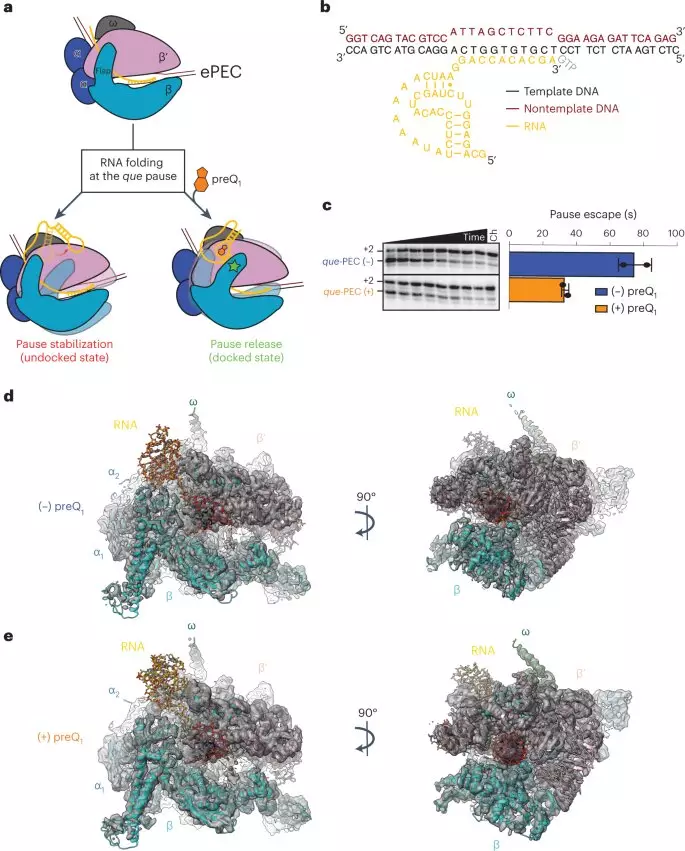
Single particle cryo-electron microscopy (cryo-EM) is a structural biology technique that can determine the structures of large protein complexes by building 3D models from millions of 2D images of particles frozen in different orientations, revealing structures with molecular details that provide functional insights. The structural information from single particle cryo-EM confirmed the Walter lab’s earlier findings and revealed a specific change in the riboswitch’s shape never seen before. When the preQ1 molecule binds, the riboswitch twists to communicate to the RNAP to continue transcription.
These observations were rationalized and validated through a collaborative effort with former U-M scientist Aaron Frank, an expert in computational modeling of RNAs. With detailed 3D models in hand, the U-M team has a more precise understanding of how this riboswitch regulates transcriptional pausing.
Potential for New Antibiotics
The new findings related to riboswitches could be used to develop new antibiotics that target these critical parts of bacterial life and avert the crisis of multidrug-resistant bacteria. According to Adrien Chauvier, a Walter lab scientist and riboswitch expert, visualizing how preQ1 regulates transcriptional pausing is like finding a needle in a haystack. The team’s approach of combining biochemistry, structural biology, and computational modeling has led to important and novel discoveries.
The University of Michigan’s multidisciplinary collaboration between three labs with different areas of expertise has led to a greater understanding of riboswitches and their role in the regulation of transcriptional pausing. The investments the university has made in cryo-EM and RNA biology in recent years have made these discoveries possible. The combination of these research efforts could lead to the development of new antibiotics that target riboswitches, preventing a global health crisis caused by multidrug-resistant bacteria.


Leave a Reply